
About
Background: Species traits are increasingly recognised as critical for understanding organisms, communities and ecosystems and their responses to global change. The GLAD project was initiated at the International Union for the Study of Social Insects conference in 2006.
Initially, the GLAD covered only species richness and abundance data for entire assemblages, but it was expanded in 2010 to include data on species abundances from local assemblages and from 2011 to include data on species traits.
At the launch of this website in October 2015, the GLAD database included more than 82910 trait values and 2212 species and 1818 georeferenced local assemblages of ants. Parr et al. (2017) details the types of data available, process through which data is acquired and summarises the data available at the time of publication.
Aim: Create a comprehensive web-based geo-referenced database on:
- local-scale species assemblages and
- species traits
in order to better understand the structure and function of species, assemblages and ecosystems and responses to global change.
Data sharing: Data is available to users for analysis and publication via the TERMS OF USE. Data collection methods and structure are outlined in Figure 2 and in Parr et al. (in review).
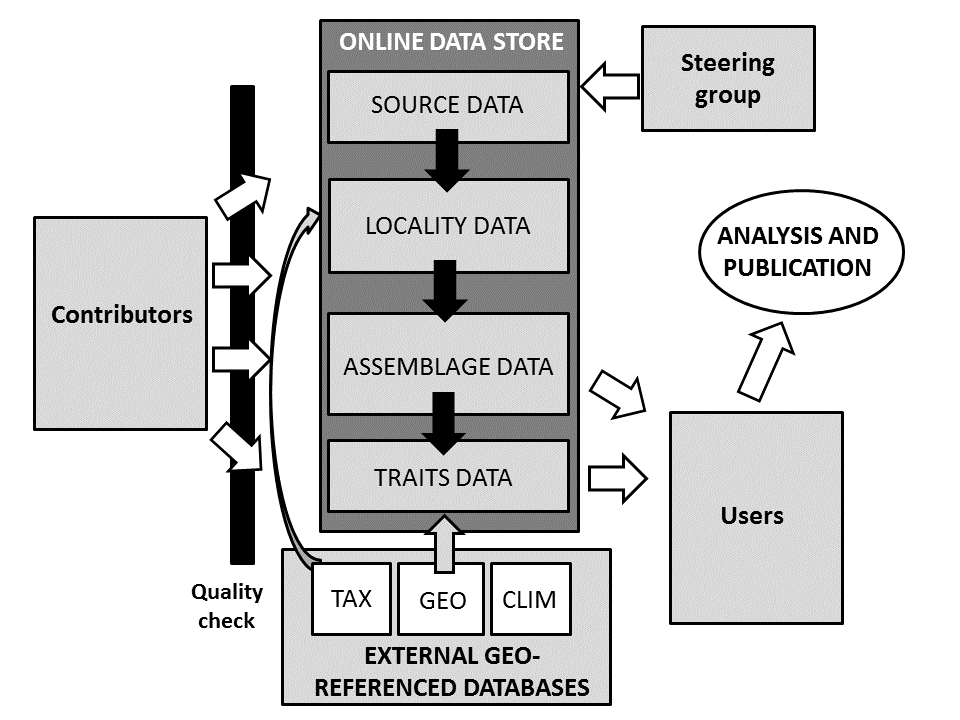
Figure 2. The Global Ant Database functions via the contribution of data which is uploaded online or emailed to the Database Managers. All data must have source and locality information. Data quality and formatting is checked prior to integration into the database. Contributors can include species abundance (assemblage data) and species trait data, but it is not necessary to have both. External geo-referenced databases (e.g., taxonomic (TAX), geographic (GEO) or climatic (CLIM) data) can be linked to either the locality or traits data as these both include information on site location. Data within the online data store area available to users for analysis and publication via a data sharing agreement.
Why ants? Ants (Hymenoptera: Formicidae) are an ideal group to study in this way because:
- ants comprise the dominant fraction of animal biomass in most terrestrial communities;
- ants perform a range of important ecosystem functions (Folgarait 1998);
- ant sampling uses standard methods of pitfall and Winkler trapping, making inter-site comparisons possible;
- ant workers are abundant, so likely to be trapped if present;
- ants are diverse, but more manageable than many other insect groups;
- data on ant morphology is obtainable from museums worldwide;
- ants are well described relative to other easily trapped groups;
- tests of relationships between ant traits and trait function are easily performed across multiple species at small scales and trait-function relationships are likely to be globally consistent (Gibb & Parr 2010);
- a robust phylogeny of ants to the genus level exists (Moreau et al. 2006), based on nuclear and mitochondrial genes (rather than morphological traits);
- new research shows strong relationships between functionally important morphological traits, habitat complexity and disturbance for ants (Bihn et al. 2010, Gibb & Parr 2010, Silva & Brandão 2010).
Support: The GLAD is hosted by La Trobe University in Melbourne, Australia. It has been funded by the Australian Research Council and La Trobe University.
Citation
Please cite the following:
Assemblage data
Gibb, H., Sanders, N.J., Dunn, R.R., Watson, S., Photakis, M., Abril, S., Andersen, A.N., Angulo, E., Armbrecht, I., Arnan, X., Baccaro, F.B., Bishop, T.R., Boulay, R., Castracani, C., Del Toro, I., Delsinne, T., Diaz, M., Donoso, D.A., Enríquez, M.L., Fayle, T.M., Feener, D.H., Fitzpatrick, M.C., Gómez, C., Grasso, D.A., Groc, S., Heterick, B., Hoffmann, B.D., Lach, L., Lattke, J., Leponce, M., Lessard, J-P., Longino, J., Lucky, A., Majer, J., Menke, S.B., Mezger, D., Mori, A., Munyai, T.C., Paknia, O., Pearce-Duvet, J., Pfeiffer, M., Philpott, S.M., de Souza, J.L.P., Tista, M., Vasconcelos, H.L., Vonshak, M., Parr, C.L. (2015) Climate mediates the effects of disturbance on ant assemblage structure. Proceedings of the Royal Society of London B: Biological Sciences. 282: 20150418. http://rspb.royalsocietypublishing.org/content/282/1808/20150418.full
AND
Gibb, H., Sanders N.J., Dunn, R.R., Abril, S., Angulo, E., Armbrecht, I., Arnan, X., Baccaro, F.B., Bishop, T.R., Boulay, R., Castracani, C., Del Toro, I., Delsinne, T., Donoso, D., Enríquez, M.L., Fayle , T. M., Fitzpatrick, M., Gómez, C., Groc, S., Gunawardene, N., Heterick, B., Hoffmann, B., Járdán, C., Klimes, P., Lach, L., Laeger, T., Lattke, J., Leponce, M., Longino, J., Lucky, A., Luke, S., Menke, S., Mezger, D., Moses, J., Munyai, T.C., Pacheco, R., Pak Nia, O., Pearce-Duvet, J., Pfeiffer, M., Philpott, S., Resasco, J., Silva, L.S.R., Silva, R.R., Sorger, D. M., Souza, J., Suarez, A., Tista, M., Vasconcelos, H.L., Vonshak, M., Yates, M & Parr, C. L. (2015) The Global Ants Database http://globalants.org/
Traits data
Parr, C. L., Dunn, R. R., Sanders, N. J., Weiser, M. D., Photakis, M. , Bishop, T. R., Fitzpatrick, M. C., Arnan, X., Baccaro, F. , Brandão, C. R., Chick, L. , Donoso, D. A., Fayle, T. M., Gómez, C. , Grossman, B. , Munyai, T. C., Pacheco, R. , Retana, J. , Robinson, A. , Sagata, K. , Silva, R. R., Tista, M. , Vasconcelos, H. , Yates, M. , Gibb, H. (2017), GlobalAnts: a new database on the geography of ant traits (Hymenoptera: Formicidae). Insect Conservation & Diversity, 10: 5-20. doi:10.1111/icad.12211
AND
Gibb, H., Sanders N.J., Dunn, R.R., Arnan, X., Baccaro, F., Bishop, T. R., Chick, L., Donoso, D., Fayle , T. M., Glasier, J., Gómez, C., Grossman, B., Munyai, T. C., Pacheco, R., Plowman, N., Retana, J., Sagata, K., Sanders, N. J., Silva, L. S. R, Silva, R.R., Tista, M., Vasconcelos, H. L. & Parr, C. L. (2015) The Global Ants Database http://globalants.org/